CRYSTAL STRUCTURE OF AMINOTRANSFERASE PRK07036 FROM Rhodobacter sphaeroides
Patskovsky, Y., Toro, R., Freeman, J., Do, J., Sauder, J.M., Burley, S.K., Almo, S.C.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Aminotransferase | 476 | Cereibacter sphaeroides 2.4.1 | Mutation(s): 0 Gene Names: RHOS4_35670, RSP_3534 EC: 2.6.1 | 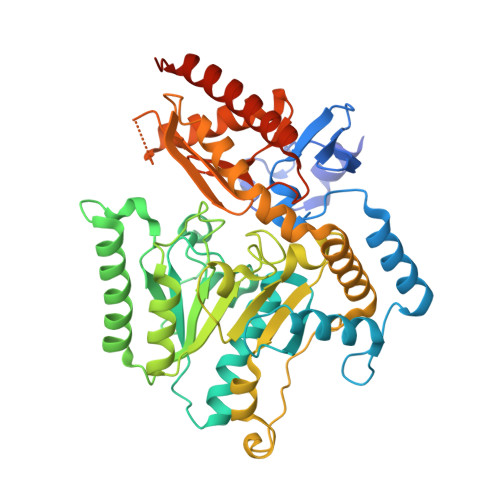 | |
UniProt | |||||
Find proteins for Q3IWE9 (Cereibacter sphaeroides (strain ATCC 17023 / DSM 158 / JCM 6121 / CCUG 31486 / LMG 2827 / NBRC 12203 / NCIMB 8253 / ATH 2.4.1.)) Explore Q3IWE9 Go to UniProtKB: Q3IWE9 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q3IWE9 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PLP Query on PLP | C [auth A], D [auth B] | PYRIDOXAL-5'-PHOSPHATE C8 H10 N O6 P NGVDGCNFYWLIFO-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 55.129 | α = 103.79 |
| b = 63.28 | β = 100.76 |
| c = 69.752 | γ = 107.08 |
| Software Name | Purpose |
|---|---|
| SHELXD | phasing |
| REFMAC | refinement |
| DENZO | data reduction |
| HKL-2000 | data scaling |